1st semester

DC1 FRANCISCO JAVIER SUAREZ LOPEZ (GENAR)
September 2025: Arrival at the host laboratory and initial familiarization with ongoing project, datasets, pipelines and literature. Training in the bioinformatic tools and workflows used for data processing and downstream analyses.
October 2025: In-depth study and implementation of software tools for the visualization and interpretation of repeat patterns, length and mosaicism in DM2.
November 2025: Integration and comparison of multi-platform LRS sequencing data to support, validate and refine tandem repeat characterization in DM2.
December 2025: Continued LRS data analysis to characterize DM2 repeat expansions, including initiation of comprehensive methylation analyses, data visualization and refinement of analytical tools.
January 2026: Evaluation and benchmarking of tools for methylation data extraction, analysis, and visualization. Refinement of analytical workflows to ensure accurate detection and interpretation of methylation patterns in DM2. First ENTRY-DM network-wide training event in Paris.
February 2026: Analyses of methylation profile and consolidation of results.
DC11 vasilina gavra (INSERM)
October 2025: Completed lab safety training and onboarding procedure. Reviewed key literature related to the project topic. Learned basic laboratory techniques and standard operating protocols. Became familiar with equipment, software, and data management systems. Held initial meetings with supervisor to define research objectives and prioritize needs.
November 2025: Completed training in sterile techniques and cell culture handling. Became familiar with myoblast cell lines, including routine culture and maintenance. Performed amplification of cell lines and established frozen stocks for future use. Conducted bibliography research and participated in brainstorming to evaluate and select the most suitable extracellular vesicle (EV) isolation strategy
December 2025: Received training on in vitro differentiation of myoblasts into myotubes. Performed screening of differentiation conditions, including plating density, incubation time, supplement addition, and coating requirements. Compared multiple experimental conditions to determine the optimal protocol for myogenic differentiation. Evaluated the differentiation profile of different myoblast cell lines. Received training in the Jess protocol for protein quantification and standardized EV-related antibodies using DM1 cell lysates. Conducted preliminary testing of a candidate biomarker previously studied in the lab using ELISA, based on the mouse DM1 model.
January 2026: Initiated collaboration with the EV platform at Institut Curie. Expanded and scaled up myoblast cultures to obtain sufficient material for extracellular vesicle (EV) production. Conducted bibliography research on size exclusion chromatography (SEC) as a method for EV isolation. Attended the first ENTRY-DM network-wide training event (INSERM, Paris), and presented my PhD project during the meeting.
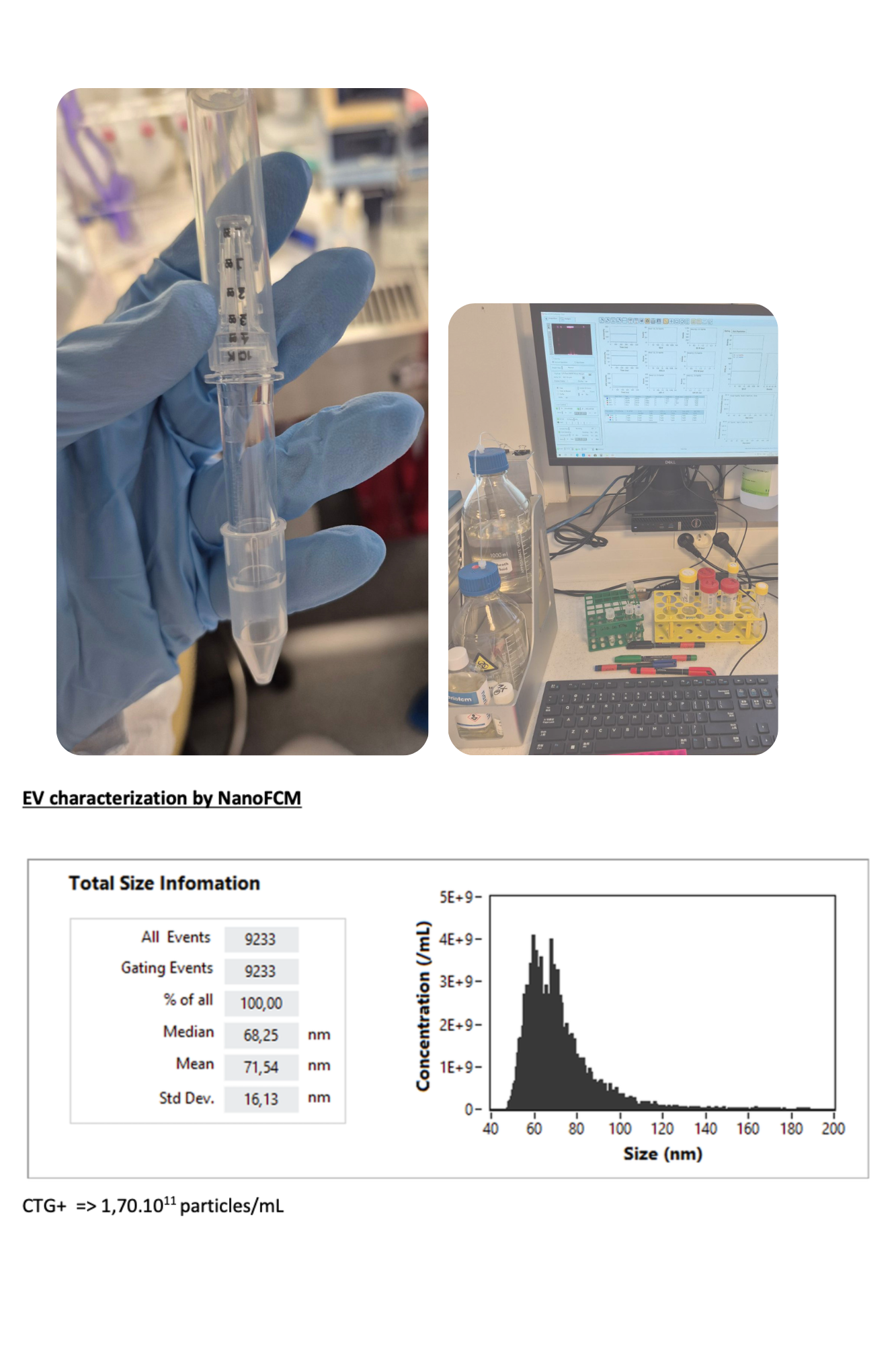
February 2026: Initiated collaboration with the Proteomics platform at Institut Curie. Received training and performed a first trial of EV isolation from differentiated myotubes. Conducted EV characterization and quantification using NanoFCM (size distribution and concentration analysis). Participated in troubleshooting and brainstorming sessions to optimize experimental conditions. Trained in RNA extraction from EVs using the miRNeasy Micro Kit. Analyzed and interpreted results from the first EV isolation experiment.
March 2026: Performed troubleshooting of myoblast differentiation to improve experimental consistency. Received training in RNA extraction and conducted splicing experiments to assess alternative splicing of genes relevant to DM1 in cell models. Continued evaluation of a candidate biomarker in a mouse model, focusing on its sensitivity to therapeutic response. Conducted a second EV isolation experiment. Performed initial proteomics analysis of EV samples to assess quality. Carried out Western blot analysis (Jess system) to evaluate EV marker enrichment and sample purity.
DC5 sofia gaspari (RUMC)
October 2025: I arrived at my host institution and started to know the spaces and complete the required documentation.
November 2025: I studied DM literature to better understand the state of the art of the topic of my project, and I outlined the research questions and the set up for my PhD trajectory. I also set up initial experiments.
December 2025: I started performing cell culture and RNA FISH experiments, to gain confidence with the techniques and to start answering the research questions defined before. I also concluded some documentation. With my lab mates we started planning for the IDMC15 conference in May 2026.
January 2026: I started planning the new year experiments and activities. I attended the kick off meeting of ENTRY-DM in Paris where I presented my research project to the PI and DC team.
February 2026: I performed the experiments previously planned (FISH), and I designed and planned new ones. I designed and ordered new probes. I gave a presentation for the journal club of the department. With the other candidates we chose the new logo for the ENTRY-DM consortium.
March 2026: I continued with my experiments and started to design the poster for IDMC15 in May. I also participated in the first online ENTRY-DM training event.

DC7 ANTON SIMÕES (RUMC)
October 2025: Settling in, getting acquainted with the patient derived iPSCs, and the project. Learning from previous members and results.
November 2025: Starting a new stable cell line for patient 3 to be used to generate iNeurons.
December 2025: First test differentiations into iNeurons, to test new stabilized cell line. Learning, and differentiating patient derived iPSCs into human astrocytes.
January 2026: Restarting cell culture. Expanding patient derived and CRISPR-edited (Ctrl) cell lines, to generate a backup of vials to be used throughout the PhD.
February 2026: Tested differentiation efficiency of newly generated patient-derived stable cell line compared to its isogenic control in which the repeat expansion was excised via CRISPR-editing. Starting Astrocyte Differentiation to be used later for human-human co-cultures.
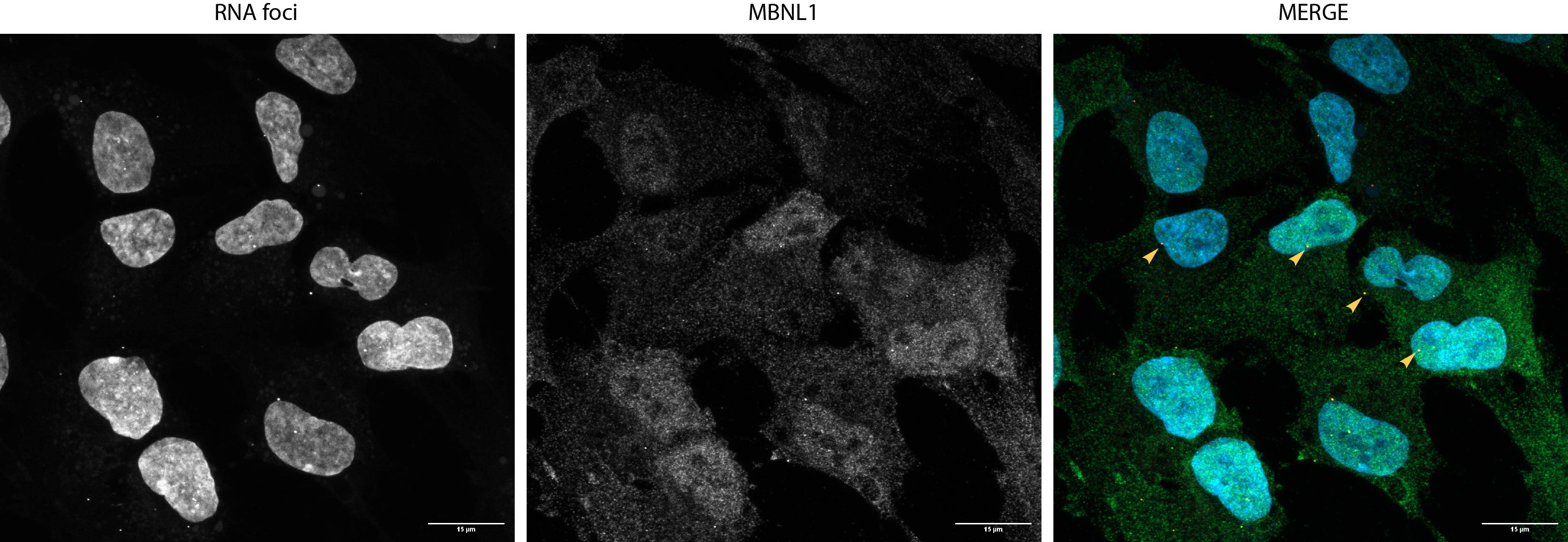
March 2026: Started first human-human co-culture with DM1 iNeurons and Astrocytes and their respective controls. Imaged first iNeurons and Astrocytes, to see if I can replicate previous results (about MBNL2 sequestration etc.). Started to collect RNA from human Astrocytes and iNeurons at different DIVs to get insights into which genes are alternatively spliced.
The image shows 28 day old iNeurons. They are generated from induced pluripotent stem cells (iPSCs) that come from a patient with Myotonic Dystrophy Type 1. Using a staining technique called FISH-IFA, multiple structures can be visualized. In Cyan we have the Nucleus. Red stains the RNA repeat expansions that show up as small puncta inside of the nucleus. And in Grey we have MAP2 which is a cytoskeleton marker that specifically labels neurons. The large nucleus visible in the top left corner belongs to a rat astrocyte. Co-culture with astrocytes is essential to support neuronal health and function, we are currently working on a protocol to incorporate patient-derived human astrocytes.

DC6 ÀNGELA NATIVIDAD I MATEU (UVEG)
October 2025: I was introduced to the laboratory members and all the facilities. I also began the process of obtaining my lab coat, university access card for entry to all buildings and facilities, and parking access. I initiated and completed the occupational risk prevention training course, compulsory for research staff at the University of Valencia.
November 2025: I had my first presentation at the lab meeting, where I presented the work carried out during my master’s thesis. Weekly meetings with my supervisors to plan future experiments. I began writing a systematic review together with DC9 on the latest advances in pipeline development for therapies for myotonic dystrophy type 1
December 2025: I started writing the Career Development Plan. We continued working on the manuscript on DM1 therapeutic pipelines. Weekly meetings with my supervisors to plan future experiments. I started my training in cellular culture. Christmas break.
January 2026: I registered for the International Myotonic Dystrophy Consortium (IDMC-15) and submitted two abstracts that got accepted. I reviewed my Career Development Plan with my supervisors and completed it. Following completion, we submitted it. I prepared and sent myotonic dystrophy type 2 (DM2) biopsy samples for RNA-seq analysis at both the transcriptome level (messenger RNA) and the miRNome level (small RNAs). I had an online meeting with my supervisor for my first secondment at Adam Mickiewicz University (AMU) in Poznań, Poland. We continued working on the manuscript on DM1 therapeutic pipelines. Weekly meetings with my supervisors to plan future experiments. We held the first ENTRY-DM kick-off meeting with all PhD candidates in Paris, France.
February 2026: We completed the systematic review on DM1 pipelines together with DC9, and the manuscript has been submitted to a journal and is currently under review. I carried out my first two outreach activities within the ENTRY-DM doctoral network. Weekly meetings with my supervisors to plan future experiments. We initiated the writing of a second manuscript with the DM2 biopsies data results from the RNA-seq. I started and completed the first transversal activity of the Doctoral School from the University of Valencia: “Ethics of scientific research”
March 2026: I started and completed two more transversal activities at the Doctoral School of the University of Valencia: “Transfer of Research Results” and “Scientific Writing.” Weekly meetings with my supervisors to plan future experiments. We received the plasmid construct from AMU in Poznań, Poland and prepared and validated it for use in the laboratory. Fallas break (national festivity that only takes place in Valencia). I had two online training sessions within the ENTRY-DM network with the following topics: Research ethics and Open Science practices, FAIR science and data sharing, Gender equality: implementation of the gender-mainstreaming strategy and PI career path. I initiated pilot experiments in DM1 immortalized myoblasts to validate the TGF-β pathway as a regulator of the dysregulated miRNAs miR-23b and miR-218. These experiments were delayed due to mycoplasma contamination in the cell line, which was successfully eliminated, allowing the work to resume. In parallel, I conducted a complementary experiment using siRNA against SMAD3 in HeLa cells to further assess the role of the TGF-β pathway in regulating miR-23b and miR-218.

DC8 MIREIA GROMAZ (AMU)
October 2025: I conducted an extensive literature review of scientific papers related to my project and other publications from the host laboratory in order to understand the scientific background and ongoing research.
November 2025: I began performing preliminary experiments to become familiar with the laboratory techniques and workflows used in the group. During this time, I also started attending the courses included in the doctoral training program.
December 2025: I continued conducting preliminary experiments to further develop my technical skills and optimize experimental conditions. I also regularly attended weekly Friday seminars from the Faculty of Biology of Adama Mickiewicz University.
January 2026: I presented my preliminary results during lab meetings and received feedback from the research group. I also travelled to Paris for one week as part of the MSCA program, where I attended seminars and training activities.
February 2026: I reviewed literature on antisense oligonucleotide (ASO) design and explored different platforms and tools for designing ASOs. In parallel, I researched immortalized myoblast cell lines suitable for testing new ASOs.
March 2026: I am currently performing experiments in several cell lines using previously tested ASOs and analyzing their effectiveness using qPCR.






